Sample definition
This section covers the sample definition step.
Sample creation involves the association of files to sample names and the addition of optional sample metadata related to the patient (e.g., phenotypes).
Once the files are uploaded, the samples can be created and defined.
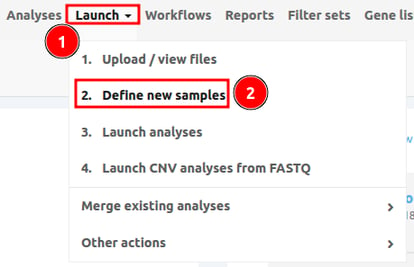
Go to Launch > 2. Define new samples as shown below.
This step is required before conducting an analysis.
Please note that SV and/or Repeat Expansion VCFs do not need to be defined separately, as they can be added during the sample definition of the main analysis.
DNA Samples
Sample definition can be performed for FASTQ and VCF files.
-
Uploaded FASTQ files are automatically recognized based on the Illumina, MGI or Element Bioscience naming conventions, and paired-end reads (which have the same filename prefix) will be grouped appropriately.
-
Uploaded VCF files are associated with a suggested sample name.
The files uploaded by the current user are shown first, followed by the files uploaded by other users. All files are sorted by their upload date.
To create samples, select the sample with the suggested sample name and follow the instructions. To select all samples simultaneously, click the “Select” option at the top of the table.

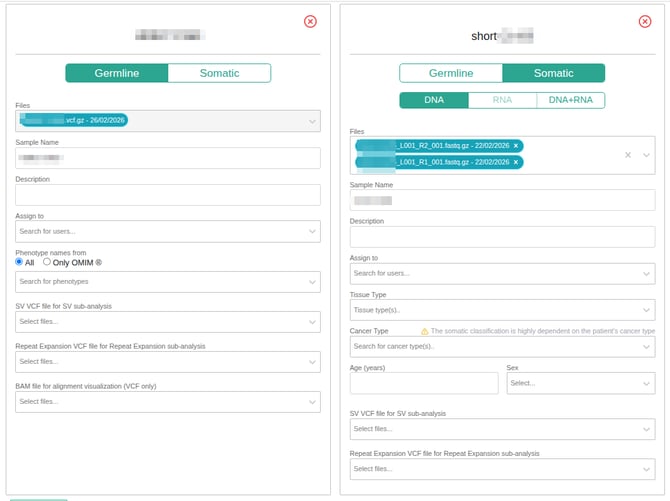
The following parameters can then be modified:
- Sample type: Click on the “germline” or “somatic” box to select the appropriate sample type.
- Files: Remove or add files.
- Sample name: Edit the suggested name of the sample here. Do not include any information about patient identity in the sample ID.
- Description: Add a description of the sample. Do not include any information about patient identity in the sample description.
- SV VCF file for SV sub-analysis (optional): Along with the sample VCF/FASTQ files, you can select a VCF file containing SVs (CNVs and/or any other type of SV). The SV results will be displayed as a sub-analysis that is accessible from the main analysis of the sample.
- Repeat Expansion VCF file for Repeat Expansion sub-analysis (optional): You can add a VCF file containing repeat expansion (RE) variants for annotation. The RE results will be displayed as a sub-analysis accessible from the main analysis.
- BAM file for alignment visualization (VCF samples only) (optional): You can add a BAM file, which will facilitate visualization of read alignments and access to IGV or JBrowse from the variant table. This file will only be used for visualization purposes, not for variant calling.
Depending on whether the sample type is germline or somatic, the relevant optional fields below will be shown:
Germline samples:
- Phenotypes: To select phenotypes of interest, begin typing a term, and then select from the list of associated phenotypes that appear. Phenotypes shown in the dropdown list may be limited based on the database source. Selecting “All” will display terms from HPO, MONDO, and OMIM® databases, while selecting “Only OMIM®” will display terms solely from the OMIM® database. These terms can be used to create a gene list and filter the variant analysis for the genes matching the selected phenotypes. For more details, see Phenotype Matching.
Somatic DNA samples:
- Tissue type (optional): Specify the tissue type of your sample.
- Cancer type (optional): Specify the cancer type for the sample.
- Age (optional): Specify the individual age of the sample (in years).
- Sex (optional): Specify the individual's sex.
⚠️ Please note that, while all the somatic sample fields are optional, provision of this information is encouraged, as it will be considered when annotating the variant with information from cancer databases.
Fusion Samples
Sample definition can be performed for fusion-RNA TSV only or DNA files (FASTQ/VCF) plus RNA TSV/CSV, as well.
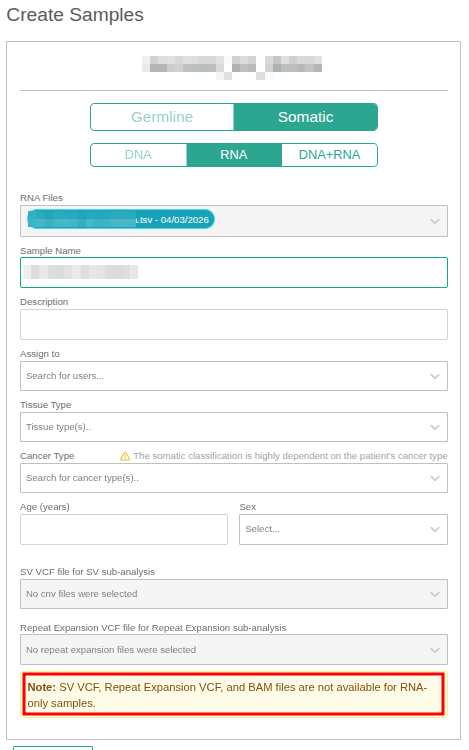
Fusion-RNA sample definition
Upon entering the sample definition page, the Analysis Type is automatically set to Somatic if a fusion TSV/CSV file was selected in the previous step. To proceed with an RNA-only sample, the RNA option must be selected and the remaining fields completed. Structural Variant (SV) and Repeat Expansion (RE) analyses are not available when running a fusion RNA-only analysis.
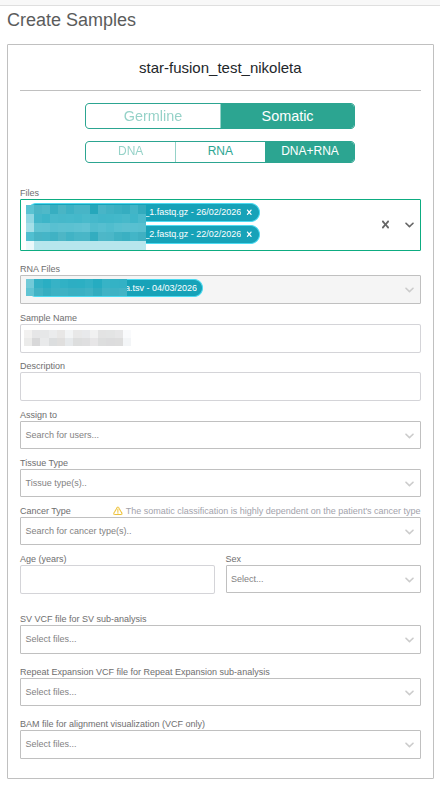
DNA + RNA sample definition
To define a combined DNA and RNA sample, the DNA + RNA option must be selected. The DNA files chosen in the previous step will automatically appear in the Files field. To add a fusion RNA file, open the drop-down menu in the RNA File for Fusion Analysis field, where the available fusion RNA files are listed. Select the appropriate file to include it in the sample definition.
As soon as the sample is created, you can now perform a new analysis. More information about the Launch analysis procedure can be found in the following documents:
Finally VarSome Clinical provides a pipeline to annotate SVs from VCF files. More information about the SV annotation can be found in the document SV annotation (from VCF).