Report files to download
From the Variant Table page of VarSome Clinical at the  drop-down menu under option "Downloads" you may find several files ready to be downloaded:
drop-down menu under option "Downloads" you may find several files ready to be downloaded:
- VCF File: A compressed (*.vcf.gz) VCF file will be downloaded with the results of the variant calling.
- For sub-analysis (gene list analysis and algorithmic filters) there is the new option to “Generate VCF” that contains only the filtered variants.
- BAM File: The bam file (the sample’s reads aligned against the reference genome) used in the analysis. For multi sample analyses you will find the BAM (and BAI) files for each component sample.
- BAI File: The index file for the above BAM file. This is a companion file for your previous BAM file, which doesn't contain any sequence data but acts as an external table of contents allowing a computational tool to navigate in the BAM file and locate specific parts.
- QC control file: A quality control report with information about the Sequence technology, Read alignment results, Regions reported, Coverage, Number of identified variants by class, Summary for Germline Variant Classification Rules and Number of SNV found in coding regions. You can download the QC report in PDF or Word format. For further details you can go here.
From the "Analysis actions" menu you can also find the related options to download several files for further investigation:
- Coding coverage report: Excel document containing details of the coverage of the coding regions included in the analysed gene list.
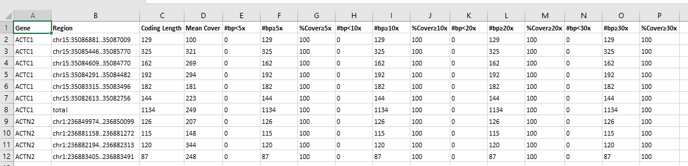
- Coverage report for targeted regions: Excel document containing details of the coverage of the assay’s target regions.
- Region list coverage report:Excel document containing details of the coverage of public/custom defined regions.
- FastQC report: A quality control report for high throughput sequence data. This file can only be viewed. You can find detailed information here.