Sequencing assay selection
When launching a new analysis without workflows, you will be asked to select the "Assay", which refers to the capture or amplicon kit used for sequencing your sample.
This selection is mandatory when starting the analysis from FASTQ files, as it defines the target regions for alignment and variant calling.
If the analysis starts from VCF files, selecting an assay is optional, since the assay information is not used in the variant annotation.

Your assay is not in the list?
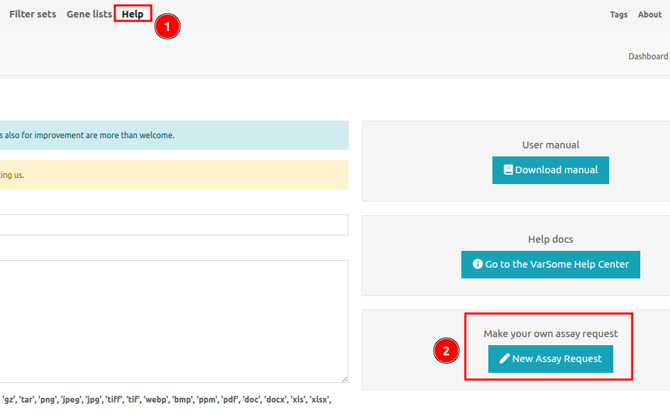
If your assay is not listed, you can request us to add it by navigating to Help > New Assay Request and filling all the required details before submitting the request.
You can find more information in the following two documents:
What is the assay information used for?
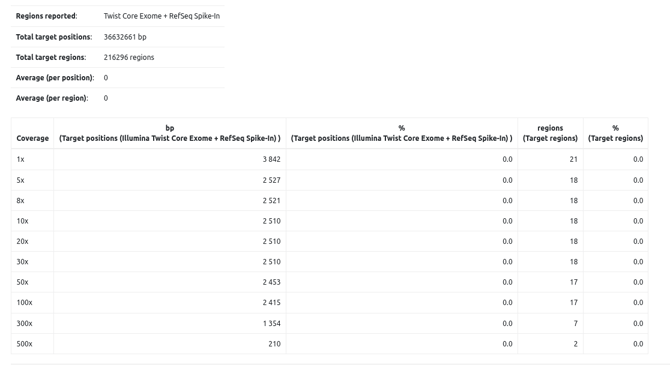
The assay defines the genomic coordinates targeted during sequencing. This information is especially relevant when launching an analysis from FASTQ generated by a sequencing experiment that is not whole-exome or whole-genome (e.g. a gene panel). In these cases, variant calling is performed in targeted mode, meaning VarSome Clinical will only call variants within the assay’s defined regions.
Additionally, assay’s information is used to calculate alignment statistics and coverage of the targeted regions. The statistics are used to generate the QC report.